How do you manage to survey a population of a small, endangered species that lives underground for most of the year? Capture-and-release, radio tracking, and other traditional methods of surveying populations might be too expensive and time-consuming. It’s an interesting conundrum that wildlife biologists and veterinarians face when trying to track animals like the Eastern Tiger Salamander (Ambystoma tigrinum tigrinum).
Like most endangered species, the Eastern Tiger Salamander has felt the impacts of anthropogenic activity. Habitat fragmentation, habitat loss, and pesticide use have threatened its livelihood and decimated its populations. Despite having a pretty broad distribution throughout the United States, it is listed as an endangered species within the state of New York. Population surveys are a bit outdated, and because Eastern Tiger Salamanders are small and spend most of their lives underground, it’s difficult to get a better understanding of changes in geographic distribution. Thankfully we can address this challenge using quantitative PCR (qPCR) to detect the DNA that these animals shed into their environments. Detecting this environmental DNA (eDNA) has been used to track movements of invasive and endangered species. Now, conservationists want to use qPCR assays to detect the presence of various species from environmental samples. The Wildlife Health Lab at Cornell University’s Animal Health Diagnostic Center has been working on developing qPCR assays for many aquatic and semiaquatic species. This past summer, I joined the lab to develop a prototype assay for the Eastern Tiger Salamander.
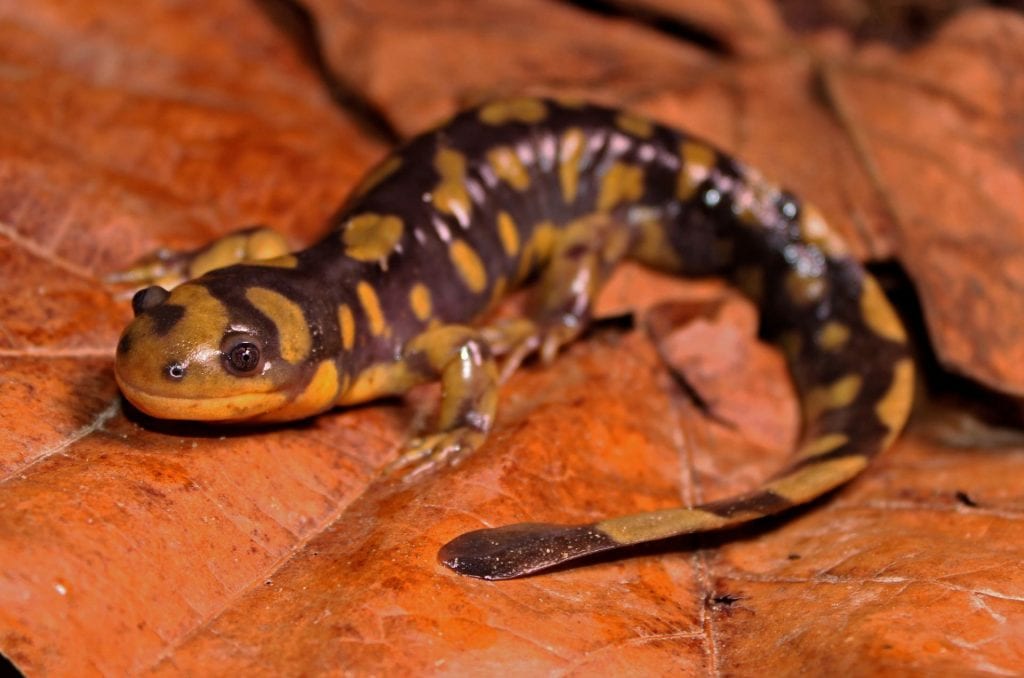

The Eastern Tiger Salamander (Ambystoma tigrinum). Credit: Peter Paplanus
Although these salamanders spend most of their time underground, they migrate once a year to breeding ponds to find mates and lay eggs. For the brief period of time they’re in the water, they shed cells from their gastrointestinal tract and secretions. DNA within these salamander cells can be collected, extracted, and detected with qPCR.
The way qPCR works can be a little complicated. In a nutshell, the goal is to massively replicate a part of the animal’s DNA sequence, attach a fluorescent probe to it, and then have a machine measure the amount of DNA that was amplified. Successful amplification means that there was a level of fluorescence that exceeded the reaction’s threshold for significance. In order for the test to work, I had to identify a part of the Eastern Tiger Salamander’s DNA sequence that isn’t found in any other living organism on the planet. Sounds like a daunting task, but thankfully the National Center for Biotechnology and Information has developed the program BLAST and the database GenBank to make it easier to identify these unique parts of the genome. While some students interested in wildlife conservation spent their time exploring the outdoors, I spent my time shifting through the Eastern Tiger Salamander’s mitochondrial genome.
Once my points of interest were discovered, I designed primer-probe combinations and then ordered the assay that was most biologically stable and least likely to react with other species’ genomes. Finally! After months of digging through thousands of nucleotide base pairs, it was finally time to test the assay. The Wildlife Health Lab had collected samples of tissue from the Eastern Tiger Salamander and some of its closest relatives. On a leap of faith we tested all these tissues against my qPCR assay.
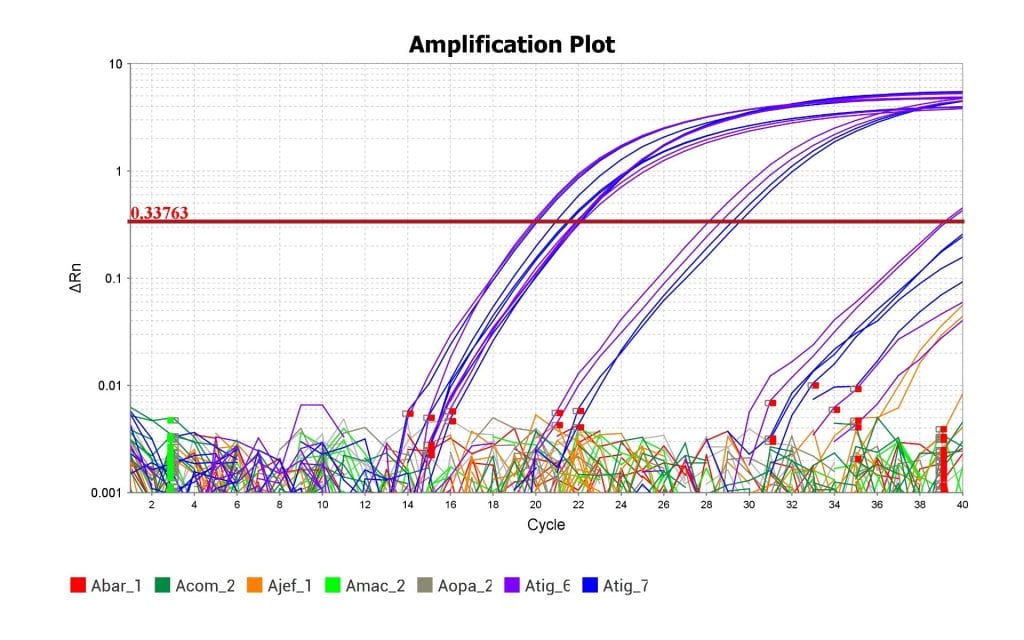
Above is the amplification plot with the Eastern Tiger Salamander’s tissue samples in blue and purple. The plot shows that the qPCR assay was able to amplify the DNA of the Eastern Tiger Salamander but not the DNA of related species. The assay was therefore successful.
As you can see from the amplification plot, this test was able to successfully replicate and detect DNA from the Eastern Tiger Salamander’s genome and not replicate the DNA from closely related species. Success! Now we can use this prototype assay to start optimizing it for efficiency and sensitivity. Hopefully, my lab can start testing it against samples of water collected from areas where we think the Eastern Tiger Salamander might be hiding.
Although we got a working prototype going, there is still much more that needs to be done. My hope is that we’ll one day have a test that can help us collect field data on the geographic distribution for many amphibian species. Once the data is generated, biologists and veterinarians can begin to map out plans for the protection of our amphibian diversity along the Northeastern coast.
Joining the Wildlife Health Lab’s eDNA project was a fantastic way to be exposed to novel diagnostic techniques and new approaches to wildlife conservation. I look forward to working with them more in the future to continue exploring other unique avenues of conservation research. Hopefully the diagnostic tool that I have helped developed will elucidate the whereabouts of one of our rare salamander species and contribute to its preservation. If its application is successful, we will be one step closer to preserving all of our local salamander biodiversity.
Beck Turcios, class of 2021, is a veterinary student at Cornell University who is interested in wildlife pathology. She graduated from the University of Notre Dame in 2016 where she studied biology and philosophy. Her experiences and interests revolve around conservation education and captive wildlife management.



