Aging is accompanied by dynamic changes in the regulatory mechanisms of gene regulation, including transcription factor activities, histone modifications and overall chromatin structure, RNA processing, and others. Importantly, research in model organisms point to a causal relationship between altered gene regulation mechanisms and consequences on the aging process and longevity.
1) Specific chromatin regulators in stress response and longevity determination
- We previously identified the transcriptional co-regulator HCF-1 and the H3K4me3 reader SET-26 to play key roles in longevity determination. Our recent findings demonstrated that SET-26 and HCF-1 are functional partners in gene regulation and longevity determination. Mechanistically, SET-26 is recruited to chromatin via the binding of its PHD domain to the histone mark H3K4me3. SET-26 then recruits HCF-1 to chromatin via its highly conserved HCF-1 interacting motif. The two factors have highly congruent genomic binding patterns and co-regulate the mRNA expression of a subset of their binding targets. We are excited to further investigate the spatial and temporal regulations of HCF-1 and SET-26 and their associated protein complexes. Both HCF-1 and SET-26 are highly conserved and delineating their functional relationship will inform the understanding of their mammalian counterparts.
2) Global analysis of the chromatin patterns that inform stress and aging biology
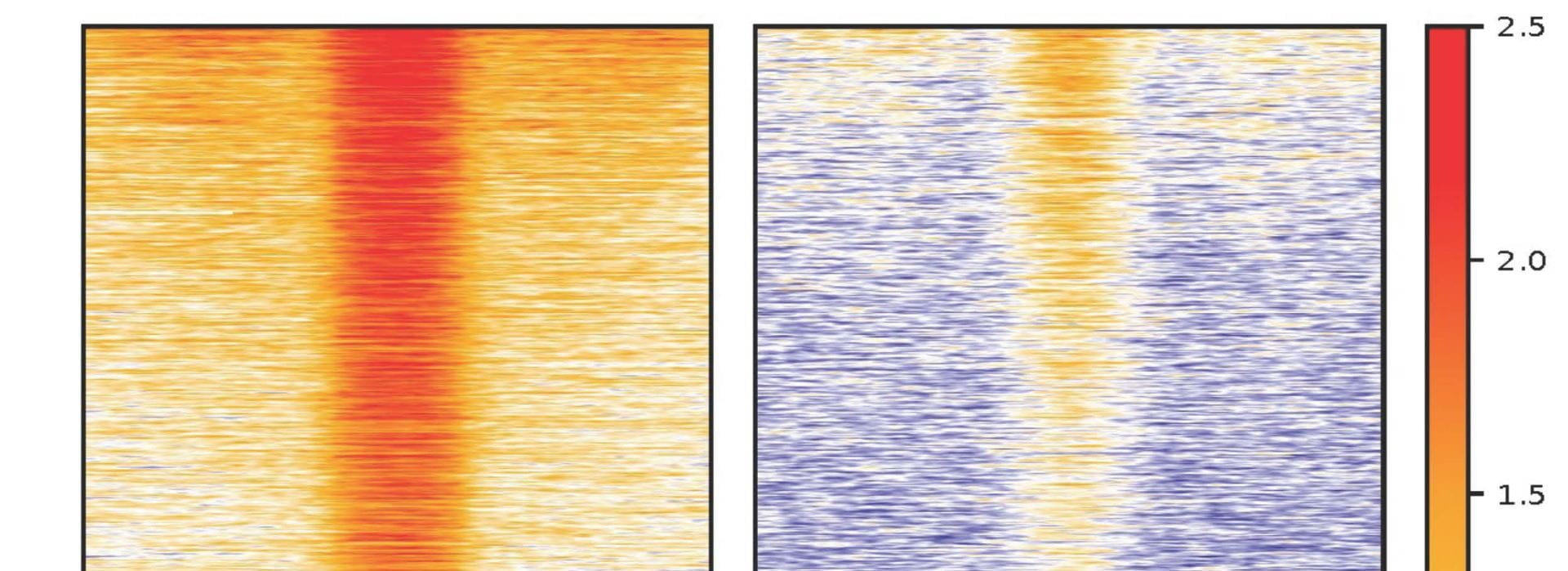
- We have profiled the genome-wide patterns of several key histone modifications through aging. Our findings have provided a much-needed framework for integrating emerging functional data to understand how global chromatin structure modulates aging and longevity in C. elegans. These genomic data also provided new hypotheses relevant to aging biology that we are excited to test, including accumulation of de novo H3K9me3 in genic regions with aging. H3K9me3 is well-known to be associated with heterochromatin, and how their accumulation in genic regions might impact aging remains to be investigated. We additionally revealed specific activation of a family of retrotransposon with aging and will be excited to further explore its functional significance.
- Stress hormesis, referring to the observation that brief exposure to mild stresses result in beneficial protective effects when re-exposed to stressors later in life. The molecular mechanisms of hormesis are not well understood. We have developed a robust regime which enables genomic strategies to probe the mechanistic underpinnings of stress hormesis.