Members
Principal Investigator:
John Lis
Laboratory Manager:
Janis K. Werner
Research Associate:
Abdullah Ozer, Senior Research Associate
 I am a Research Associate and I have been in the Lis Lab since 2007. I got my Bachelors degree in Molecular Biology and Genetics from Bilkent University, Turkey, and my Ph.D. in Molecular Biophysics from UT Southwestern Medical Center at Dallas, Texas. In my graduate study working with Richard K. Bruick in Department of Biochemistry at UTSW, I characterized the function of ING4 protein in regulation of mammalian Hypoxia Response Pathway, the primary cellular response mechanism for oxygen deprivation. I have a general interest in developing novel high-throughput technologies that enable us, in ways that were not possible before, to disrupt, image, and characterize biological processes particularly in transcriptional regulation in live cells. Since I have joined the Lis Lab, I have been involved in many projects including the improvement of RNA Aptamer selection and characterization methods and their implamantaton for selection and characterization of RNA aptamers to a number of proteins including many transcription factors, and imaging of proteins and their post-translational modifications with RNA aptamers in living cells. Recently, I have been working on developing novel and improved methods for determination of 4D nuclear architecture and characterization of enhancers with high resolution and high sensitivity in mammalian cells. Also in the last three years I have been involved in a challenging project called “Raising Irem and Harun”, serving as the father (trying hard to be a good one).
I am a Research Associate and I have been in the Lis Lab since 2007. I got my Bachelors degree in Molecular Biology and Genetics from Bilkent University, Turkey, and my Ph.D. in Molecular Biophysics from UT Southwestern Medical Center at Dallas, Texas. In my graduate study working with Richard K. Bruick in Department of Biochemistry at UTSW, I characterized the function of ING4 protein in regulation of mammalian Hypoxia Response Pathway, the primary cellular response mechanism for oxygen deprivation. I have a general interest in developing novel high-throughput technologies that enable us, in ways that were not possible before, to disrupt, image, and characterize biological processes particularly in transcriptional regulation in live cells. Since I have joined the Lis Lab, I have been involved in many projects including the improvement of RNA Aptamer selection and characterization methods and their implamantaton for selection and characterization of RNA aptamers to a number of proteins including many transcription factors, and imaging of proteins and their post-translational modifications with RNA aptamers in living cells. Recently, I have been working on developing novel and improved methods for determination of 4D nuclear architecture and characterization of enhancers with high resolution and high sensitivity in mammalian cells. Also in the last three years I have been involved in a challenging project called “Raising Irem and Harun”, serving as the father (trying hard to be a good one).
Postdocs:
 Sagar Shah (Joint with Haiyuan Yu’s Lab)
Sagar Shah (Joint with Haiyuan Yu’s Lab)
Due to the diversity and complexity of signaling networks in both physiological and disease states, I am interested in understanding the mechanisms that drive transcriptional programs and its regulation. During my doctoral studies in Biomedical Engineering at Johns Hopkins University School of Medicine and subsequent work at the Mayo Clinic, my research focused on studying transcriptional dysregulation in cancer by delineating novel YAP-mediated signaling networks that drive aggressive behavior for prognostic and therapeutic purposes. Currently, as a joint postdoctoral fellow in the Lis and Yu labs, I am characterizing the active transcriptional regulatory element landscape of cancer and studying its regulation to understand fundamental mechanisms underlying gene expression.
Gopal Chovatiya (Joint with Tudorita Tumbar’s lab)
Gopal received his PhD in Life Sciences from the Advanced Centre for Treatment, Research and Education in Cancer (ACTREC), India. He is fascinated by the robust and dynamic regeneration potential of adult stem cells that reside in skin epithelium, and developments of genomic technologies for interrogating transcriptional regulation. As a shared postdoc between the Lis lab and the Tumbar lab, he is currently working on developing a new methodology that would not only enable studying finest transcriptional regulation in vivo, but also potentially unravel novel regulatory mechanisms for stepwise gene expression during stem cell differentiation.
Graduate Students:

Miliarys Quinones Barreto
(Genetics, Genomics and Development)
I’m from Moca, Puerto Rico. I obtained my Bachelor of Science (BS) in Biology with an emphasis in Genetics at the University of Puerto Rico at Aguadilla. My research as an undergraduate student focused on gene regulation and epigenetics. Currently, I’m a PhD student in the Graduate Field of Genetics, Genomics and Development (GGD). I’m interested in investigating the release mechanisms for productive elongation of promoter-proximal paused RNA Polymerase II and the role of this step on the regulation of gene transcription.
Brent Basso (Joint with Charles Danko’s Lab)
(Computational Biology)
James Borovilas (BioPhysics)
 Caralyn Gonzales
Caralyn Gonzales
(Chemistry and Chemical Biology)
Originally from Eagle, Idaho, I was awarded my B.S in Chemical Biology from University of California, Berkeley and am currently a graduate student in the field of Chemistry and Chemical Biology. My work focuses on the glucocorticoid stress response as a concurrent backdrop to numerous diseases. I combine rapid in vivo perturbations with genome-wide measurements to study how the mechanisms of transcriptional regulation under stressful conditions interplay with cancer pathogenesis. Outside of lab, I enjoy hiking, volunteering, reading, traveling, and any DIY project that can hold my attention.
 Yiyang Jing (Joint with Haiyuan Yu’s Lab)
Yiyang Jing (Joint with Haiyuan Yu’s Lab)
(Biochemistry, Molecular and Cell Biology)
Investigating the Mechanism of Enhancer-Promoter Specificity in Humans
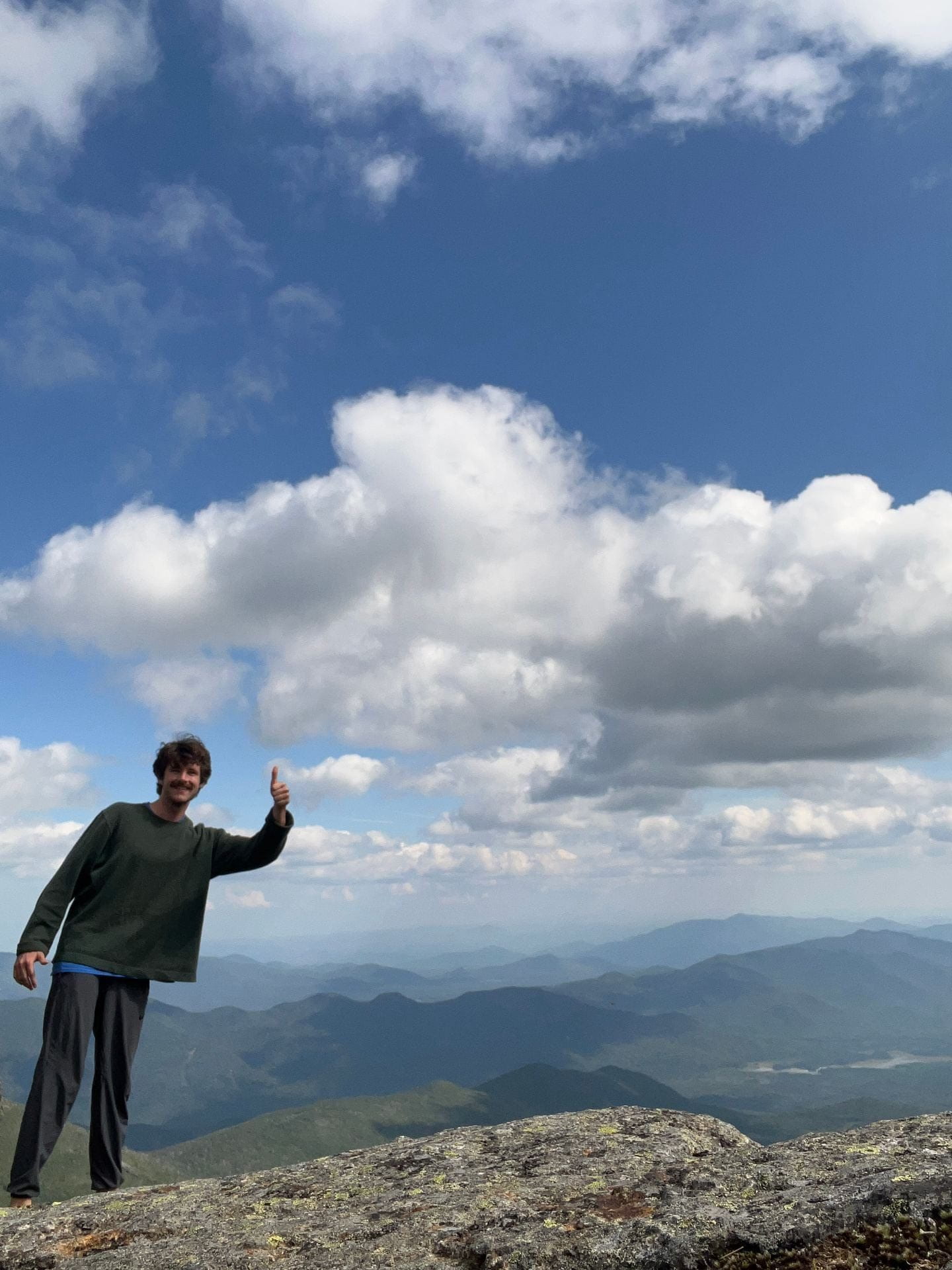 Jaret Lieberth
Jaret Lieberth
(Genetics, Genomics and Development)
I grew up outside of Cleveland, OH, in a town called Aurora. From there I went south to do my undergraduate at the University of Mississippi, where I picked up my first genetics/molecular biology experience in the lab of Joshua Bloomekatz. My project there involved finding the functional tissue location of platelet-derived growth factor receptor alpha (pdgfra) during early cardiogenesis using zebrafish. I graduated a bit unsure if I wanted to pursue an MD or a PhD, so decided to head west for a technician job in the lab of Nathan Clark at the University of Utah to help me decide. There my work involved characterizing a large set of putative cis-regulatory elements presumed to play a role in eye development, also in zebrafish. It was this project that sparked my interest in the basics of transcription, especially how noncoding elements can work to influence it. Now, as a PhD student in the Genetics, Genomics and Development program I’m co-advised by Cedric Feschotte and John Lis. My work currently (not in zebrafish this time) focuses on utilizing nascent transcription mapping as a method for enhancing causative SNP identification in disease, as well as investigating the role of proteins involved in the early innate immune response. When I’m not in the lab you’ll find me reading, hiking Ithaca’s lovely trails, or playing volleyball (or tennis, basketball – anything I can find a partner for).
 Philip Versluis
Philip Versluis
Zhou Zhou
__________________________________________________________________________________________________________
Undergraduates
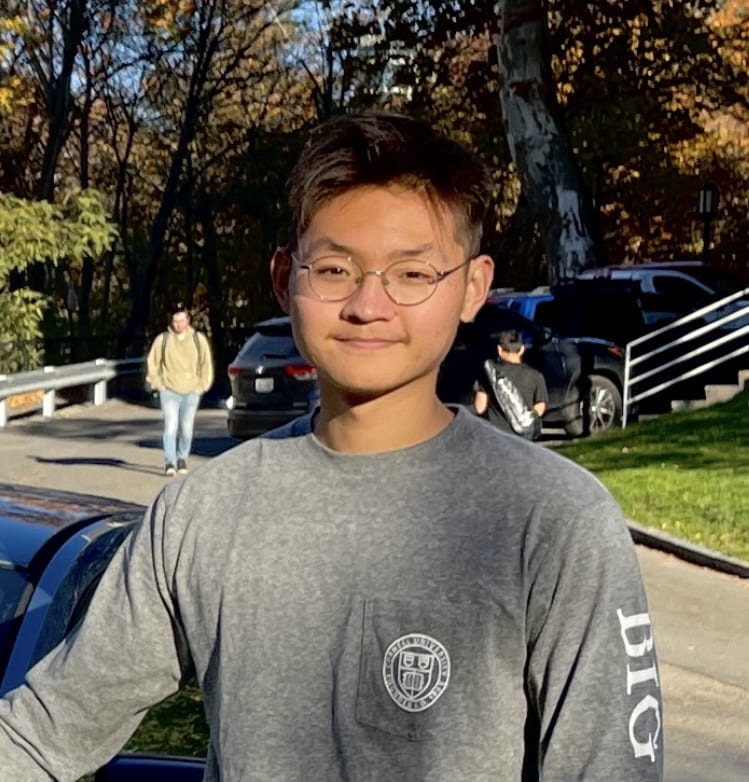 Albert LiAlbert is an undergraduate student majoring in Biological Sciences with a concentration in Molecular and Cell biology. His current study in the lab involves robotic-assisted generation of tagged mammalian cell lines, for the study of specific transcription-related proteins.
Albert LiAlbert is an undergraduate student majoring in Biological Sciences with a concentration in Molecular and Cell biology. His current study in the lab involves robotic-assisted generation of tagged mammalian cell lines, for the study of specific transcription-related proteins.


Leave a Reply